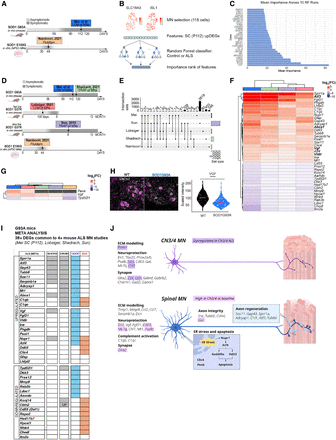
Data set comparison and feature selection show a common importance for Vgf in ALS. (A) Overview of the comparison between two data sets: the Namboori et al. (2021) (human) and SC (P112; mouse) data sets. (B) Schematic of the random forest–based machine learning approach for identifying key ALS-relevant genes using upregulated DEGs in SC (P112; n cells selected in Namboori et al. data set: 115). (C) Feature importance ranking from the random forest classifier, highlighting VGF as the most predictive marker. (D) Overview of cross-data set DEG comparison, including work of Shadrach et al. (2021), Lobsiger et al. (2007), Sun et al. (2015), and Namboori et al. (2021). (E) Upset plot showing DEGs shared across ALS data sets. Likelihood ratio test, Benjamini–Hochberg adjusted P-value < 0.05. (F) Heatmap of the 39 common DEGs identified across all mouse studies between SC at P112, Shadrach et al. (2021), Sun et al. (2015), and Lobsiger et al. (2007). Bolded gene names are also predictive in the random forest disease classifier. (G) Heatmap showing common DEGs between all data sets. Colors represent log2(FC); black squares indicate that that gene did not result as a DEG for the relative data set. (H) Representative immunofluorescence image of SC at P112 and showing the expression of VGF protein in VAChT-positive cells. Scale bar, 30 µm. Welch's two-sample t-test P ≤ 2.2 × 10−16. (I) Heatmap displays the 39 ALS-induced DEGs (F) common to four mutant SOD1 MN studies (Mei, Lobsiger, Shadrach, and Sun). Genes are categorized as upregulated or downregulated in ALS MNs. The presence of each gene in different data sets is indicated in columns MEI (disease-induced DEGs shared between CN12 and SC MNs at P112 from Mei et al.) and CRUSH (sciatic nerve crush model from Shadrach et al. 2021), as well as if the regulation is considered “GOOD” or “BAD.” (J) Summary of gene expression and regulation in CN3/4 and spinal (SC) MNs in healthy animals and SOD1-ALS. (Top) CN3/4-enriched genes are involved in ECM remodeling, neuroprotection, and synapse function, with those regulated in ALS highlighted in purple. (Bottom) Gene expression dysregulated in spinal MNs in SOD1-ALS is involved in ECM remodeling, neuroprotection, complement activation, synapse function, axon integrity, ER stress and apoptosis, and axon regeneration. Genes normally enriched in healthy CN3/4 MNs compared with spinal MNs that are induced in spinal MNs with ALS include the neuroprotective genes En1, Fgf21, Mt-Tq, and Pvalb and ECM-related genes.











