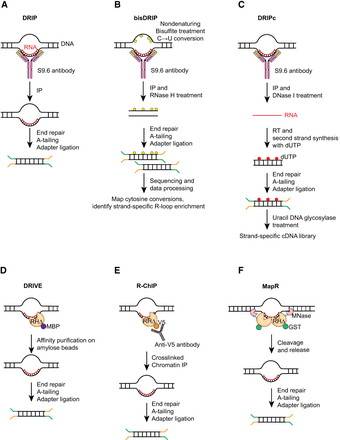
Schematic overviews of key methodologies used for mapping R-loops. (A) In DRIP, fragmented genomic DNA is incubated with the S9.6 antibody to immunoprecipitate DNA–RNA hybrids for sequencing. (B) bisDRIP combines DRIP with nondenaturing bisulfite treatment to selectively deaminate ssDNA regions and convert cytosine residues into uracil. Detection of cytosine conversion events in sequenced libraries enables the identification of the ssDNA and DNA–RNA hybrid components of the R-loops in a strand-specific fashion. (C) DRIPc combines immunoprecipitation with the S9.6 antibody with DNase I treatment to selectively degrade the DNA strands, leaving the RNA component of DNA–RNA hybrids. Strand-specific cDNA library preparation allows identification of R-loop RNA. (D) In DRIVE-seq, catalytically deficient RNase H (RHΔ) is used to isolate DNA–RNA hybrids from fragmented genomic DNA prior to sequencing. (E) In R-ChIP, RHΔ is expressed in vivo with an epitope tag, and genomic sequences bound by RHΔ are recovered via chromatin immunoprecipitation. (F) MapR uses a purified RHΔ–MNase fusion protein to bind and cleave chromatin fragments containing R-loops, which are then purified and sequenced.











