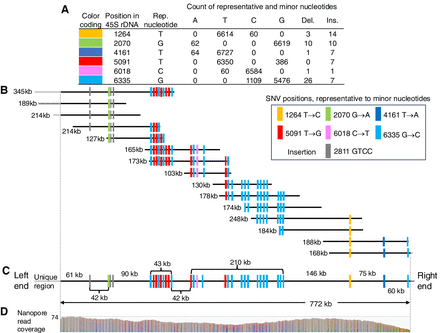
45S rDNA array at the right end of CHROMOSOME_I. (A) Single-nucleotide variants (SNVs) detected in the 45S rDNA representative repeat unit of size 7197 nt; 3322 PacBio HiFi reads were aligned to the 45S rDNA representative repeat unit. Because each HiFi read was long enough for the 45S rDNA unit to occur approximately twice, HiFi read coverage in the 45S rDNA unit averaged 6691 at 7197 positions. The table shows SNVs whose minor nucleotides are detected 60 or more times in PacBio HiFi reads. (B) Nanopore reads are aligned so that distances of adjacent SNVs and insertions within Nanopore reads are consistent between aligned reads. (C) Consensus of aligned Nanopore reads, an assembly of the 45S rDNA array with 107 units. Each position in the consensus is covered by two or more Nanopore reads shown in B. (D) Histogram displays read coverage for each position within the consensus by Nanopore reads >100 kb in length.











