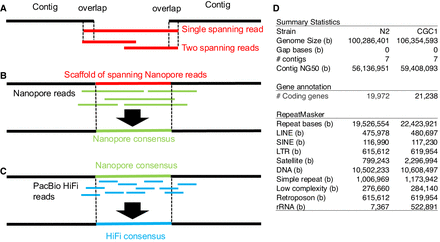
The overlap-layout-consensus approach of assembling tandem repeat regions. (A) The overlap-and-layout step finds one or more Nanopore ultralong reads that span a focal gap and lays out multiple reads properly. (B) The consensus step corrects errors in the scaffold of Nanopore reads (red), aligns other Nanopore reads (green) to the scaffold, and calculates the consensus of the aligned reads. (C) To further eliminate errors in the consensus sequence (green), HiFi reads (blue) are aligned to the consensus and used to generate the consensus of mapped HiFi reads. (D) Comparison between the N2 and CGC1 assemblies. (#) The number of contigs, coding genes, and so on. The N2 genome assembly version is WBPS19 WBcel235 (GCA_000002985.3) generated in December 2012.











