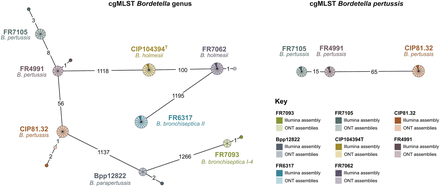
Minimum spanning trees of Bordetella pertussis and other Bordetella species investigated in this work (computed with GrapeTree). Core genome multilocus sequence typing (cgMLST) profiles used for pairwise comparisons in these trees were generated from (1) Illumina genome assemblies (dark triangles) and (2) Oxford Nanopore Technology (ONT) genome assemblies (including all assemblies from different simulated coverage depths), whose raw data were basecalled with Dorado SUP v0.9.0 (lighter colors). The tree on the left was generated from cgMLST profiles of the cgMLST_genus scheme, whereas the tree on the right from cgMLST profiles of the cgMLST_pertussis scheme (uniquely applied to B. pertussis genomes). Edges numbers indicate allelic distances between entries (edge length: log-scale).











