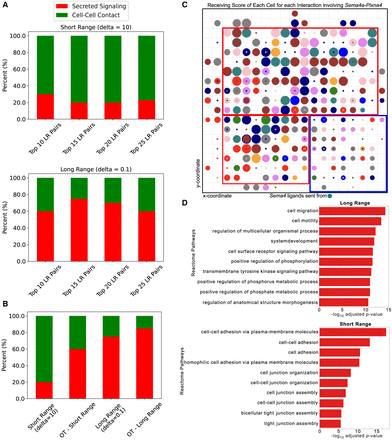
CellAgentChat identifies long-range and short-range interactions in the adult mouse cortex data set. (A) Bar plot illustrates the percentage of interactions found by CellAgentChat annotated by CellChat's database as either “cell–cell contact” (short-range) or “secreted signaling” (long-range) with a delta (δ) value set to 10 (top) and 0.1 (bottom). (B) Bar plot illustrates the percentage of interactions found by CellAgentChat annotated by CellChat's database as either “cell–cell contact” or “secreted signaling” in comparison to an optimal transport (OT)-based approach (Liu et al. 2022). (C) CellAgentChat's animation platform depicts the cell receiving score for a “cell–cell contact” Sema4a-Plxna4 ligand–receptor pair using the short-range mode (δ = 10). The color of the cells (circles) represents the cell cluster. This animation depicts the release of Sema4a (ligand) from cells in cluster 9 (represented by turquoise cells). In the animation, cell size corresponds to the cell receiving score. Cells closer to cluster 9 cells exhibit higher CRS (within red square) than cells that are further away (within blue square), which are smaller in size. (D) Reactome pathway analysis of the receptors involved in the top 25 LR interactions in the long-range mode (top) and short-range mode (bottom).











