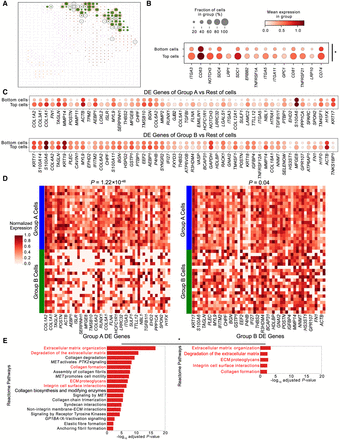
CellAgentChat facilitates inference of cellular interactions associated with individual cells in the PDAC data set. (A) CellAgentChat's animation platform enables the detection of communication strength variations in individual cancer cells. The platform highlights the cell receiving score (Methods) of Can2 cells (colored green). The top 20 cells with the highest CRS (group A) are marked by red circles and the bottom 20 cells with the lowest CRS (group B) are marked by black circles. (B) Expression analysis of receptors in the top 25 LR interactions shows significantly higher expression in group A compared to group B (Wilcoxon test one-sided; P < 0.05). (C) Mean expression of differentially expressed (DE) genes upregulated for group A spots relative to rest of the spots (top) and upregulated for group B cells relative to rest of the spots (bottom). (D) Heat maps display the DE genes for group A (left) and group B (right). The DE genes of group A cells are mainly expressed by group A and not by group B (Wilcoxon test one-sided; P < 0.001). Both groups express the DE genes of group B but are more significantly expressed by group A spots (Wilcoxon test one-sided; P < 0.05). (E) The results of the Reactome pathway analysis using group A DE genes (left) and group B DE genes (right). Red highlights indicate shared pathways.











