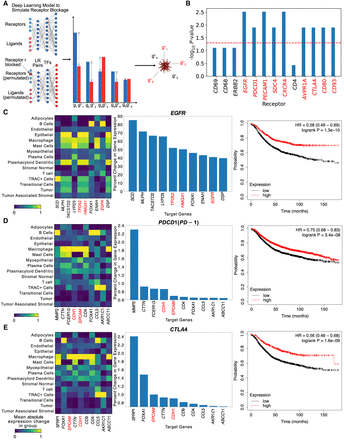
CellAgentChat enables in silico receptor blocking to identify potential receptor antagonists for therapeutics. (A) Schema of the in silico perturbation analysis using our neural network to block receptors and identify the most perturbed downstream genes. Blue bars show unperturbed predicted expression; red bars show perturbed predictions. (B) Breast cancer gene set enrichment (−log10 adjusted P-values; binomial test, FDR corrected) for significantly perturbed genes after blocking 13 selected receptors. Receptors highlighted in red are significantly enriched. (C–E) Matrix plots showing mean expression changes per cell type for EGFR, PDCD1 (PD-1), and CTLA4, respectively (left). Top 10 downstream genes with highest expression change after blocking EGFR, PDCD1 (PD-1), and CTLA4, respectively; red highlights indicate breast cancer-associated genes (middle). Survival analysis comparing high versus low receptor expression groups for EGFR, PDCD1 (PD-1), and CTLA4, respectively (right).











