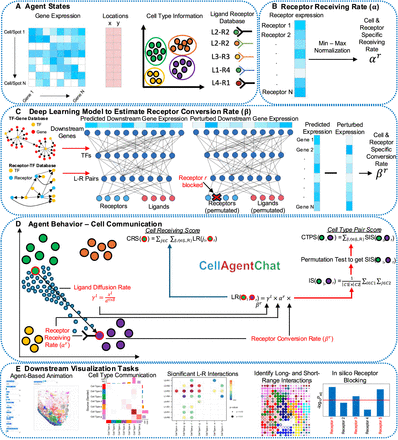
Overview of CellAgentChat. (A) CellAgentChat requires the following agent states for each cell: gene expression from scRNA-seq, cell type/cluster information, and LR database. Spatial coordinates (from spatial transcriptomics data) are optional. (B) The receptor receiving rate is the probability that an interaction is received by a receptor, calculated by non-parametric min-max normalization of receptor expression. (C) Our deep learning model inputs the gene expression of ligands and receptors, predicting the expression of all genes. The model incorporates prior knowledge of TF interactions. Receptor blocking is simulated by permuting receptor features and scaling down expression. The receptor conversion βr of the receptor is estimated from the total change of the predicted expression after the perturbation. (D) LR interaction scores depend on ligand diffusion rate (γl), receptor receiving rate (αr), receptor conversion rate (βr), and optionally spatial distance (d) between cells. Hyperparameters τ (spatial freedom, default = 2) and δ (ligand decay rate, default = 1) control interaction range. Using the LR score, we can calculate the interaction score between two clusters/cell types and the cell receiving score (CRS) for a cell. After a permutation test, significant interactions are identified and used to calculate the cell type pair score (CTPS) between two clusters. (E) CellAgentChat enables agent-based animation to visualize cell communication, identify significant LR pairs, and simulate receptor blocking to find the most perturbed downstream genes for potential therapeutic drug discovery.











