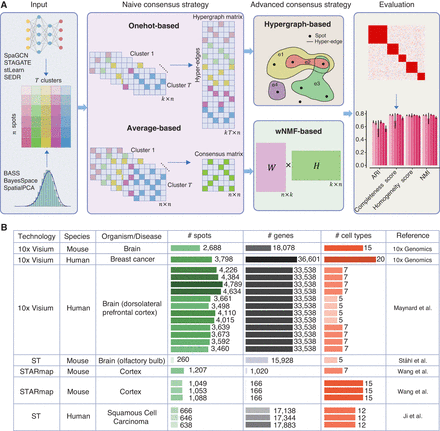
STCC architecture and evaluation data sets. (A) STCC architecture. STCC takes clustering results of seven baseline algorithms as input. It first constructs a hypergraph matrix and a consensus matrix and then executes two naive strategies (average-based, Onehot-based) and two advanced strategies (hypergraph-based, wNMF-based) to get consensus results. The ultimate clustering results are evaluated using seven benchmark metrics, namely, ARI, NMI, completeness score, homogeneity score, Calinski–Harabasz score, Davies–Bouldin score, and stability. (B) Details of seven benchmark data sets, including sequencing technologies, species, organism/disease types, numbers of spots, genes, and cell types, etc.











