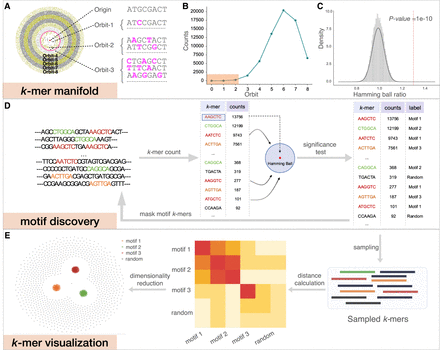
KMAP workflow. (A) Schematic illustration of the k-mer manifold for k = 8. Each point represents a unique k-mer. Orbit-i consists of k-mers with i mutations to the origin; namely, the Hamming distance from the k-mer to the origin is i. k-mers in the ith orbit are uniformly scattered in the ith ring, where each ring has an equal width. k-mers within the red circle forms the Hamming ball centered on the origin with a radius r(k) = 2. (B) k-mer counts of each orbit. The rectangle highlights the k-mer counts of orbits in the Hamming ball. (C) Null distribution of Hamming ball ratio. The histogram is generated by taking all Hamming ball ratios from a random DNA sequence of 100,000 bp. The experiment is repeated 10 times, and a Gaussian distribution is fitted to the obtained ratios with the mean fixed to one. The fitted Gaussian distribution is used as the null distribution, in which the vertical dashed line indicates the significant ratio corresponding to a P-value of 1 × 10−10. (D) The motif discovery workflow. We first count the k-mers and then test the Hamming ball centered on the top k-mer; after that, we mask all motif k-mers from the input DNA sequence and repeat the process iteratively until no motif can be found. (E) k-mer visualization algorithm; 2500 motif k-mers and 2500 random k-mers are sampled for the visualization. The Hamming distance matrix of the sampled k-mers is smoothed and further utilized for dimensionality reduction.











