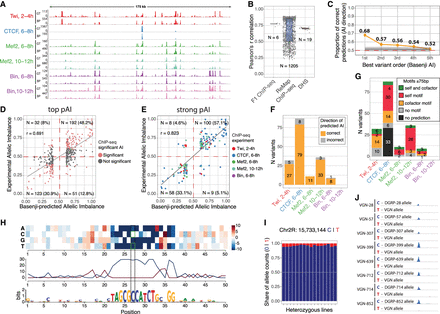
Basenji prioritizes causal variants and predicts disrupted motifs via saturation scores. (A) Ground-truth (GT) experimentally measured coverage from merged F1 crosses compared with Basenji-predicted (BP) coverage on test sequences from Chromosome 2L. (B) Pearson R correlation between GT and BP coverage on test set sequences at 1 kb resolution for all sample tracks used in training (ReMap n = 1205, DHS n = 19, F1 ChIP-seq n = 6). (C) Fraction of correct predictions (same direction of pAI and experimental allelic imbalance [AI]) for variants associated to the same peak and ranked by absolute pAI from Basenji predictions. This test includes all peaks associated with at least one variant with significant experimental allele imbalance. Gray line indicates 10,000 permutations of random variant ranking (background); light gray shadow, 2 SDs. (D) Correlation between predicted imbalance and experimentally measured imbalance for variants with the highest absolute pAI per peak (top pAI: absolute pAI > 0.1 for variants ±2.5 kb of each peak); 79.1% of Basenji predictions are in the correct direction with Pearson R = 0.691. (E) Correlation between strong Basenji pAI (absolute pAI > 0.1) and strong experimental allelic imbalance (absolute AI > 0.1); 90.2% of predictions are in the correct direction. Colors correspond to ChIP-seq samples. (F) Counts of strong predictions per ChIP-seq sample divided by correct (same direction of effect; orange) and incorrect (gray) allelic imbalance predictions. (G) Same counts as in F, with variants colored by motif predictions from saturation mutagenesis ±75 bp around the causal variants: TF's own cognate motif (red), a potential cofactor motif (orange), both the cognate motif and a potential cofactor motif (green), no motif predicted (gray), nonconverging prediction (black). (H) Saturation scores around variant Chr 2R: 15,733,144 correlating with high allelic imbalance in CTCF 6–8 h, reveals that Basenji predicts a CTCF motif that is disrupted by a C-to-T variant. (I) Proportion of allele-specific reads mapping to reference (C) or alternative (T) alleles on variant Chr 2R: 15,733,144 for CTCF 6–8 h. (J) Visualization of ChIP-seq signal separated by maternal and paternal allele on the CTCF peak affected by variant in panel I. All paternal lines harbor the C allele, whereas the maternal line has the T allele.











