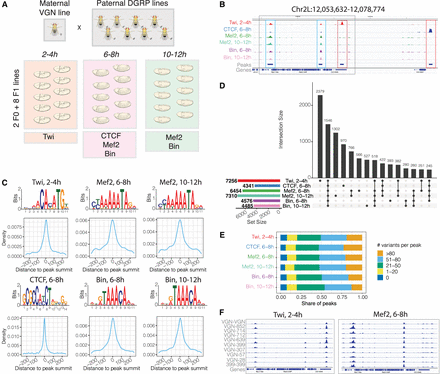
Profiling transcription factor (TF) binding in F1 embryos of Drosophila melanogaster. (A) Experimental design: F1 crosses were generated between a maternal VGN line and eight paternal DGRP lines. Staged embryos were collected at three time points (2–4 h, 6–8 h, 10–12 h). Binding of four TFs (Twi, CTCF, Mef2, and Bin) was profiled at the indicated embryonic time points, in two parental lines (VGN and DGRP399) and eight F1 lines. (B) Occupancy of all TFs in the chromosomal region Chr 2L: 12,053,632–12,078,774. For each TF, tracks were generated by averaging signal across all 10 lines at the corresponding time point. Blue boxes indicate regions cobound by multiple factors; red boxes show peaks specific to one TF. (C) De novo motifs discovered in top-1000 ChIP-seq peaks (top) and central enrichment of these motifs in full consensus peak sets of Twi at 2–4 h, CTCF at 6–8 h, Mef2 at 6–8 h, Mef2 at 10–12 h, Bin at 6–8 h, and Bin at 10–12 h. (D) Overlaps among consensus peak sets for all TFs. (E). Consensus peaks overlap for each TF with the indicated number of genetic variants (zero, one to 20, 21 to 50, 51 to 80, more than 80 variants) within 5 kb regions centered on peak summits. (F) Binding of Twist at 2–4 h (left) and Mef2 at 6–8 h (right) across the individual lines on Chr 2L: 12,054,806–12,066,910 (region marked as gray box in panel B).











