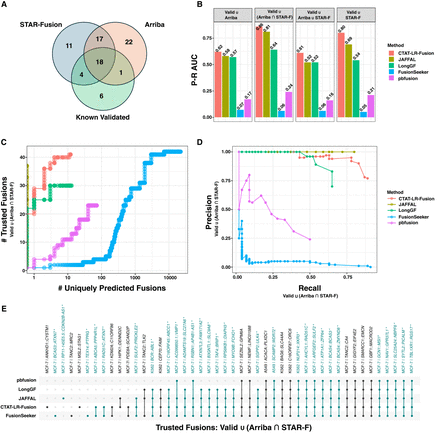
Fusion transcript detection using ONT transcriptome sequences derived from cancer cell lines. (A) Counts of trusted fusions identified from long-read isoform sequences according to known validation status and orthogonal Illumina read support as reported by STAR-Fusion or Arriba. (B) Fusion detection accuracy for methods according to alternatively defined sets of trusted fusion transcripts as truth sets and scoring uniquely predicted fusions as FPs, with accuracy measured as the P–R AUC. For C–E, trusted fusions are defined as known validated fusions or those identified by both Arriba and STAR-Fusion as having Illumina read support (see Supplemental Table S6). (C) Numbers of trusted fusions detected versus uniquely predicted fusions according to method with fusions ranked from highest to lowest fusion read support. (D) Precision–recall plot for benchmarking fusion prediction accuracy using trusted fusions as TPs and uniquely predicted fusions as FPs. (E) UpSet plot indicating which trusted fusions were identified by which methods, with trusted fusions colored teal and fusion names with asterisks.











