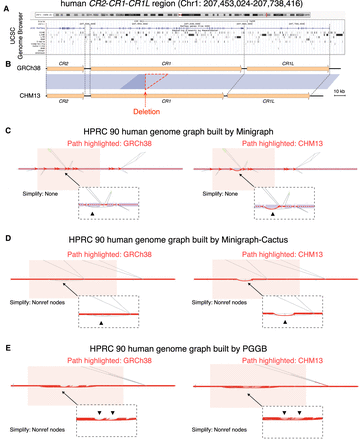
VRPG's visualization of the CR1 intragenic deletion in the HPRC human pangenome graphs. (A) The UCSC Genome Browser view for the CR2-CR1-CR1L region at Chr 1: 207,453,024–207,738,416 (GRCh38 coordinates) with annotation tracks for chromosome ideogram, gene structure, and repetitive sequence features displayed. (B) The genome sequence and synteny comparison between GRCh38 and CHM13 for the CR2-CR1-CR1L region, with the blue shades representing homologous regions shared with >98% sequence similarity. The deletion is further highlighted in the red triangle. (C–E) VRPG's visualization for the CR1 deletion in the HPRC human pangenome graphs derived from Minigraph (C), Minigraph-Cactus (D), and PGGB (E), with the genome paths of GRCh38 and CHM13 highlighted, respectively. The pink shades denote the CR1 genic region. The path differences in the deletion region are further indicated by black triangles. VRPG's squeezed layout was used for the Minigraph-Cactus and PGGB graph visualization. Nonref node simplification was applied as indicated when visualizing the Minigraph-Cactus and PGGB graphs. All other options were left with defaults.











