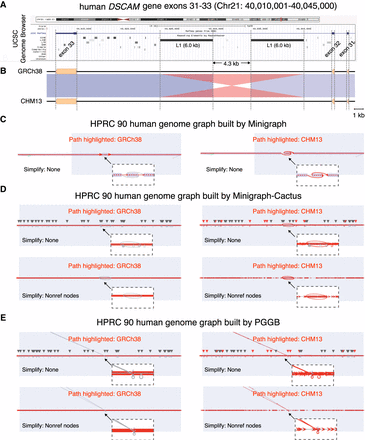
VRPG's visualization of the DSCAM intronic inversion in the HPRC human pangenome graphs. (A) The University of California, Santa Cruz (UCSC) Genome Browser view of the DSCAM exons 31–33 region at Chr 21: 40,010,001–40,045,000 (GRCh38 coordinates) with annotation tracks for chromosome ideogram, gene structure, and repetitive sequence features displayed. (B) The genome sequence and synteny comparison between GRCh38 and CHM13 for the DSCAM exons 31–33 region, with the blue (matched in same directions) and red (matched in reversed directions) shades representing homologous regions shared with >98% sequence similarity. (C–E) VRPG visualization for the DSCAM inversion in the HPRC human pangenome graphs derived from Minigraph (C), Minigraph-Cactus (D), and PGGB (E), with the assembly paths of GRCh38 (left) and CHM13 (right) highlighted, respectively. The light purple shades denote the DSCAM exons 31–33 region. The small triangles along the graph indicate the trace of genomic variants shown in the graph, with those corresponding to the highlighted path further colored in red. VRPG's squeezed layout was used when visualizing the Minigraph-Cactus and PGGB graphs. Layout simplification was applied as indicated, whereas node scaling was enabled in all cases. All other options were left with defaults.











