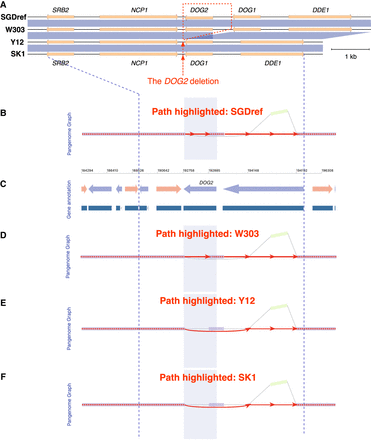
VRPG's visualization of the polymorphic DOG2 deletion in a yeast pangenome graph. (A) The genome sequence and synteny comparison among SGDref, W303, SK1, and Y12 for the DOG2 region, with the blue shades representing homologous regions shared with >97% sequence similarity. (B–F) VRPG visualization for the graph in this region (Chr VIII: 189,131–197,233, SGDref coordinates). The assembly paths of SGDref (B), W303 (D), Y12 (E), and SK1 (F) are highlighted, respectively, with the SGDref-based coordinate system and gene annotation track (C) further shown in between. The light purple shades denote the DOG2 gene region. No layout simplification or node scaling was applied in VRPG for this visualization. All other options were left with defaults.











