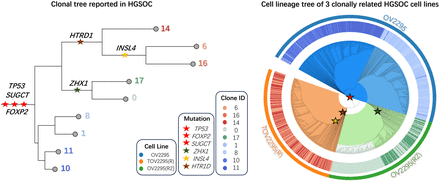
Analysis of the HGSOC data by ScisTree2. Left: The clonal tree of HGSOC with six mutated genes considered important for cancer development in the original study (Laks et al. 2019). Number at a leaf: clone ID (with a distinct color). Right: Reconstructed cell lineage tree of tumor cells (colored w.r.t. their clones) with the same genes mapped on. Outer ring: three cell lines OV2295, TOV2295(R), and OV2295(R2). Inner ring: (colored) clones from the original study. Genes marked as red stars are ancestral genes, whereas others are clade genes.











