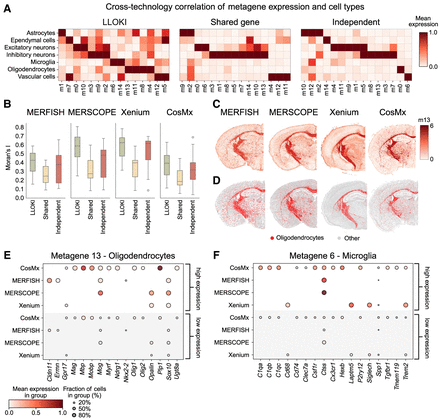
SpiceMix with LLOKI embeddings enhances cross-technology metagene discovery. (A) Cell-type enrichment of metagenes across MERFISH, MERSCOPE, CosMx, and Xenium data sets, comparing three approaches: LLOKI-based integration, the shared-gene approach (22 common genes), and independent analyses. (B) Quantitative assessment of metagene quality using Moran's I spatial autocorrelation scores, shown as box plots. (C) Spatial distribution of metagene 13 across all four technologies, showing consistent patterns using LLOKI embeddings with SpiceMix. (D) Spatial distribution of oligodendrocytes across MERFISH, MERSCOPE, Xenium, and CosMx, demonstrating the correspondence between metagene 13 and oligodendrocytes. (E) Expression levels of canonical oligodendrocyte marker genes across MERFISH, MERSCOPE, Xenium, and CosMx in cells highly expressing metagene 13 versus cells lowly expressing metagene 13. (F) Expression levels of canonical microglia marker genes across MERFISH, MERSCOPE, Xenium, and CosMx in cells highly expressing metagene 6 versus cells lowly expressing metagene 6.











