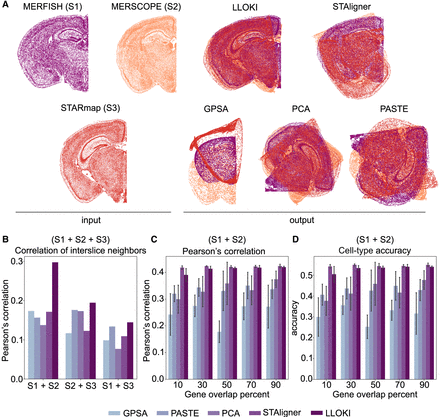
Evaluation of LLOKI's performance in spatial slice alignment. (A) In situ visualizations of three slices from the MERFISH, MERSCOPE, and STARmap data sets, along with alignment results using LLOKI and four baseline methods. (B) Alignment accuracy as measured by Pearson's correlation of interslice neighbors between each pair of slices from A. (C,D) Comparison of LLOKI and four baseline methods on alignment accuracy of the MERFISH and MERSCOPE slices as gene overlap decreases, showing results for (C) Pearson's correlation, and (D) cell-type accuracy.











