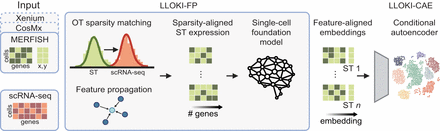
Overview of LLOKI. LLOKI performs a two-stage alignment for integrating ST data: First, it unifies the feature space across ST samples, regardless of gene panel; then, it removes technology-specific batch effects. LLOKI-FP applies optimal transport-guided feature propagation to impute missing gene expression values using spatially informed cell graphs, mitigating data sparsity and gene sensitivity differences. A single-cell foundation model then embeds the data into a unified feature space. LLOKI-CAE further integrates these embeddings across batches using a conditional autoencoder, removing technology-specific effects while preserving biological structure. The resulting embeddings minimize batch effects and support downstream tasks such as cross-technology slice alignment and spatial gene program identification.











