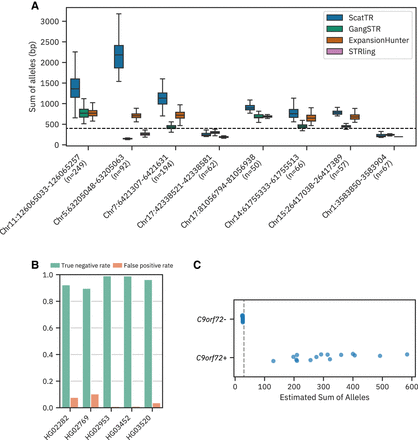
Evaluation of ScatTR on real WGS data. (A) Predicted sum of allele lengths (in base pairs) of TR expansions by ScatTR and state-of-the-art methods. The labels on the x-axis indicate the sample size for each locus. All loci have motif lengths >6 bp, except for the locus on Chromosome 5. The mean fragment length of 400 bp is shown as a horizontal line. (B) True-negative rate and false-positive rate of TR expansion calls for five samples with long-read ground truth across 171146 TR loci. Expansions are defined as exceeding the fragment length threshold of 400 bp. (C) Predicted sum of allele lengths (in base pairs) for 30 ALS patients based on short-read WGS data. The data set includes 15 samples with known C9orf72 expansions (C9orf72+) and 15 without (C9orf72−). The vertical line indicates the pathogenic cutoff of 30 repeat units. All samples were correctly classified.











