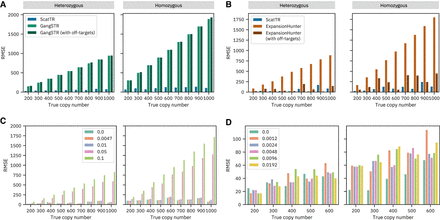
Impact of off-target regions, mutation rates, and sequencing errors on ScatTR accuracy. (A) RMSE of ScatTR compared with GangSTR, with and without off-target regions, in 108 simulated samples across different true copy numbers. Heterozygous cases are shown on the left and homozygous on the right. The simulated samples represent expansions in 12 TR loci for which GangSTR provided off-target regions, including SCA1–3, SCA6–8, SCA12, SCA17, HTT, DM1, FMR1, and C9orf72. (B) RMSE of ScatTR compared to ExpansionHunter, with and without off-target regions, in 18 simulated samples across different true copy numbers. Heterozygous cases are shown on the left and homozygous on the right. The simulated samples represent expansions in two TR loci for which ExpansionHunter provided off-target regions, including C9orf72 and FMR1. (C) RMSE of ScatTR across varying mutation rates in the repeat sequence. (D) RMSE of ScatTR across different sequencing error rates in whole-genome sequencing (WGS) samples.











