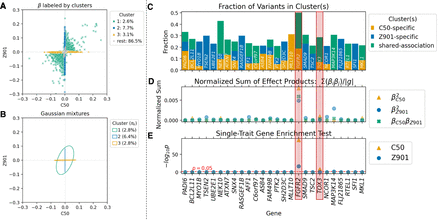
ML-MAGES’s association clustering and gene-level signals highlight genes with shared associations across two binary traits in the UK Biobank. Two traits are malignant neoplasm of breast (C50) and acquired absence of breast (Z90.1). The two genes highlighted in the figure, FGFR2 and TOX3, are those identified by Cortes et al. (2020) as showing similar biological pathways. The figure style follows that of Figure 3, E–I. (A) The bivariate clustering results based on regularized effects of two traits. (B) Inferred mixtures from bivariate clustering, shown as covariance ellipses with inferred mixing weights πk. (C) Fraction of variants in each gene that belong to each type of clusters. The genes listed have at least 10 variants and >20% of variants from one of the prioritized clusters. (D) Normalized sum of effect products of variants in each gene. (E) Single-trait enrichment test −log10(P) for each trait.











