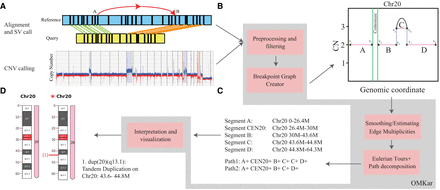
Overview of the OMKar method. (A) Input data. OMKar takes structural variant (SV) calls, copy number variation (CNV) calls, and sequence alignments as input. (B) Preprocessing and filtering. Low-confidence SVs and CNVs are removed. Chromosomes are segmented based on CNV boundaries and breakpoints, and a breakpoint graph is constructed in which vertices represent segment boundaries and edges represent segment continuity, reference adjacencies, or rearrangements. (C) Smoothing and path decomposition. Integer linear programming is used to estimate edge multiplicities while maintaining copy number consistency. Edge editing produces a Euclidean graph, and reconstructed paths are extracted using Eulerian tour and decomposition methods. (D) Interpretation and visualization. Structural variations are annotated using ISCN nomenclature; disrupted genes are identified; and results are compiled into an interactive HTML report with chromosome visualizations.











