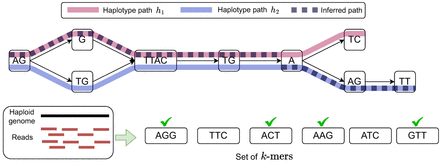
A schematic to illustrate the path inference problem. At the top, we show a haplotype-resolved pangenome graph containing two haplotype paths, h1 and h2. These paths are shown in pink and blue, respectively. An inferred path containing a single recombination is illustrated using a thick dashed line. At the bottom, we show a set of k-mers observed in the sequencing reads assuming k = 3. A k-mer is marked with a green tick if it is a substring of the sequence spelled by the inferred path AGGTTACTGAAGTT. Two k-mers (TTC and ATC) are not present in this sequence. Our optimization framework identifies an optimal inferred path with minimum cost given user-defined penalties for recombinations and for missing k-mers.











