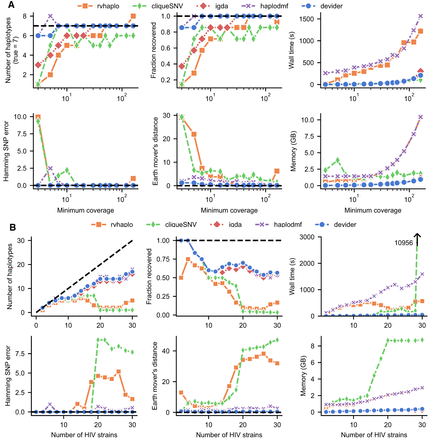
Benchmarking long-read haplotyping tools on simulated HIV-1 communities. (A) Seven HIV-1 strains from Kinloch et al. (2023) at staggered abundances (1:3:5:7:9:10:20) with simulated reads (9000 bp mean length; 95% mean accuracy). The x-axis indicates the depth of coverage for the lowest-coverage strain. (B) Two to 30 HIV-1 strains with 15×–145× uniformly random coverage and the same simulated read length/accuracy. CliqueSNV failed to complete within the default 3 h time limit for the last sample. iGDA does not output abundances, so its earth mover's distance could not be computed. Dashed black lines indicate optimal performance.











