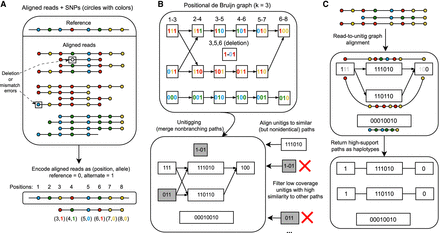
Algorithmic framework for devider. (A) Reads that are aligned to a reference are converted to a SNP representation with positional information. Sequencing errors lead to erroneous SNP encodings. (B) The SNP-encoded reads are turned into a positional de Bruijn graph (PDBG; k = 3 shown). In a PDBG, k-mers are collapsed if their alleles and their positions are identical. Errors in reads lead to spurious k-mers in the PDBG. After merging paths with in-degree and out-degree equal to one (unitigging), unitigs are aligned back to the graph to filter low-coverage, high-similarity unitigs. (C) Reads are aligned back to the filtered unitig graph to determine high-confidence walks through the graph. These paths are taken to be putative haplotypes. devider then postprocesses the haplotypes to output haplotype abundances, a base-level consensus of each haplotype, and the reads assigned to each haplotype.











