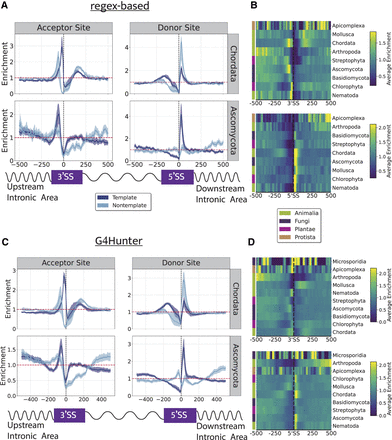
The topography of G4s relative to splice sites in eukaryotic phyla. (A) Distribution of G4s relative to 3'ss and 5'ss using G4 motifs derived from the regex algorithm for two eukaryotic phyla, Chordata and Ascomycota. (B) Heatmap showing the enrichment of G4s relative to splice sites across different phyla for both the template and the nontemplate strand combined. (C) Distribution of G4s relative to 3'ss and 5'ss using G4 motifs derived from the G4Hunter-based algorithm for two eukaryotic phyla, Chordata and Ascomycota. (D) Heatmap showing the enrichment of G4s derived from the G4Hunter algorithm relative to splice sites across different phyla for both the template and the nontemplate strand combined. Results are shown for the template and nontemplate strands separately. Confidence intervals in plots A and C represent the 2.5% lowest and 97.5% highest percentile from Monte-Carlo simulations with replacement (N = 1000).











