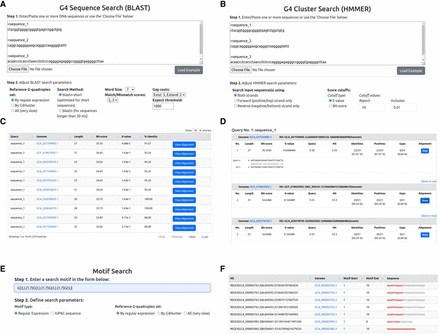
The sequence search options. (A) The G4 sequence search input page. Search options include defining the search algorithm (BLASTN), setting the word size, and defining the match/mismatch scores and gap penalties. (B) G4 cluster search. Users can choose which DNA strand to include in the search and define cutoffs for the E-value and bit-score metrics. (C,D) Example BLAST results. An overview (C) of the results presents the basic search statistics (alignment length, bit-score, E-value, and percentage sequence identity), and the detailed results (D) give the full BLAST output, including a visualization of the aligned sequences. (E,F) Input form (E) and example results (F) of the motif search functionality. Sequence motifs can be given as regular expression patterns or IUPAC sequence stretches. The results include all G4 sequences that match the submitted motif.











