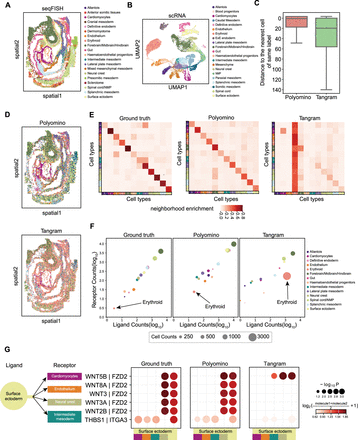
Analysis of mouse embryo data at E8.5. (A) Major tissue zones of the mouse embryo at E8.5 using seqFISH. (B) UMAP visualization of main cell types annotated by Pijuan-Sala et al. (2019). (C) Compared cell reconstructed distances of Polyomino and Tangram (x-axis). We calculated cell reconstructed distances between the mapped cells and spatial voxels of the same cell type. (D) Spatial distribution of single-cell types after reconstruction by Polyomino and Tangram. (E) Analysis of spatial neighboring cell proportions of original seqFISH, Polyomino, and Tangram. (F) Number of ligand–receptor pair analyses with original seqFISH data and Polyomino- and Tangram-reconstructed data, with circle size indicating cell counts. (G) Ligand–receptor interactions between the surface ectoderm and other cell types.











