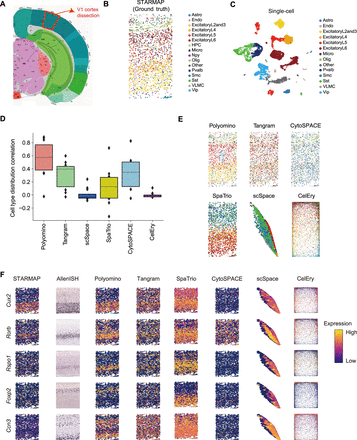
Evaluation in cerebral cortex data. (A) Tissue sections from cerebral cortex were used to generate spatial data (STARMAP) and snRNA data. (B) Spatial location of main cell types in the STARMAP data. (C) UMAP visualization of main cell types of the snRNA data. (D) Quantitative comparison of spatial reconstruction accuracy across six methods. Similarity was measured by correlating predicted cell-type distributions with the spatial ground truth. (E) Spatial mapping results from Polyomino and other representative methods. Polyomino achieves higher spatial concordance and sharper boundary definition of the cell types across cortical layers. (F) Comparison of reconstructed gene expression patterns. Polyomino better recapitulates the original spatial expression, demonstrating higher fidelity in gene-level reconstruction.











