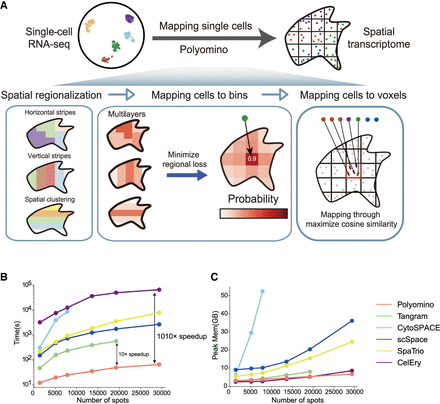
Schematic overview of Polyomino. (A) Polyomino is designed to accurately assign single-cell locations. Starting with spatial region expression formed from gridding the original ST data and single-cell seq data (left), Polyomino uses multilayer regional constraints to optimize the distribution of each cell in each grid (middle) and finally determine the precise location of cells within each grid through min cosine similarity cost (right). (B) Computation time of different methods when integrating 5000 single cells with varying voxel numbers. (C) Peak memory of different methods when integrating 5000 single cells with varying voxel numbers.











