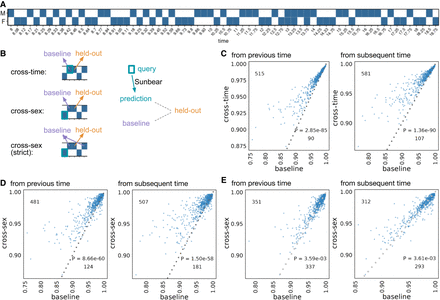
Single-cell profile inference across time and conditions. (A) Sunbear is trained on scRNA-seq profiles of whole-mouse embryos collected at alternating sexes along developmental time points (Qiu et al. 2024). (B) Sunbear is validated in three scenarios. In each scenario, one data block is held out from the training, and Sunbear is used to predict the profile of the missing block based on cells in the query block (outlined in turquoise). Sunbear's prediction is compared against the baselines using the held-out block's nearest existing measurements with the desired sex factors. The pseudobulk Pearson's correlation per major cell trajectory is calculated between the held-out profile and predictions/baselines. (C) Cross-time evaluation: query and baseline are selected either from the closest previous time point (left) or from the closest subsequent time point (right). The pseudobulk Pearson's correlation between the original held-out profile and predicted (y-axis) and baselines (x-axis) is plotted for each major cell trajectory in each held-out time point. Each dot represents a cell trajectory per held-out time point, and numbers indicate the number of dots above and below the diagonal line. P-values are calculated by a one-sided Wilcoxon rank-sum test. (D) Cross-sex prediction: similar to that in C, except query cells are selected from the opposite sex to the held-out data. (E) Cross-sex prediction: similar to that in D, except we enforce a strict baseline model by taking the mean of the previous and subsequent time point per cell trajectory.











