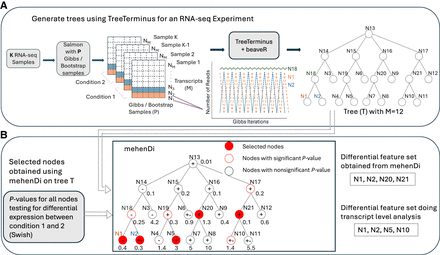
Schematic overview of mehenDi. (A) Given a set of RNA-seq samples, TreeTerminus (along with beaveR) creates a tree that encodes the uncertainty structure by using the Gibbs/Bootstrap samples generated by Salmon to compute uncertainty. (B) Given a tree, mehenDi outputs a set of selected nodes. The selected nodes are differentially expressed between the conditions of interest and can consist of transcripts and inner nodes, with no two nodes having an ancestor/descendant relationship. Using an example, with a tree having 12 leaves to demonstrate mehenDi, a highlighted node implies that it is selected by mehenDi, with a red outlined node indicating that the node has a significant P-value. The + or − on a node indicates the direction of sign change between the conditions, whereas +− indicates we are unsure about the direction. The values plotted along the nodes represent their mean inferential relative variance across the samples (meanInfRV). The meanInfRV threshold in this example is 0.4. The nodes N1 and N2 are chosen instead of node N18 because the meanInfRV of one of its children, node N2, is below the threshold of 0.4. The node N5 is chosen instead of the node N19 because the children have opposite sign changes. Node N20 is the only significant node in its branch and meets all the required criteria. Node N21 is picked because it has at least one nonsignificant child node.











