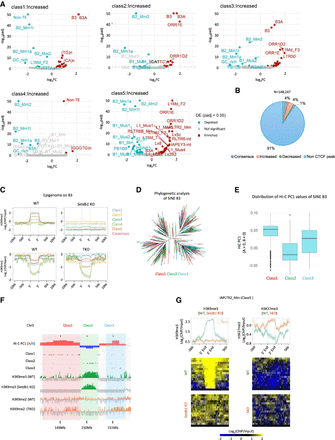
Prevention of additional CTCF binding at repetitive elements by H3K9/K27 methylation. (A) Enrichment of repeat type in increased DE peaks. The volcano plots represent the enrichment of each repeat type in DE peaks, with the x-axis showing log2(FC) and the y-axis showing −log10(adjusted P-value). Repeat types that are significantly enriched or depleted in each class compared to consensus peaks (adj. P-value < 0.01) are highlighted in red and blue, respectively. All raw data used in A were described in Supplemental Table S2. (B) Fraction of SINE B3 transposons overlapping with CTCF peaks. “Consensus,” “Increased,” “Decreased,” and “Non-CTCF peak” represent the fractions of SINE B3 copies that overlap with the “Consensus peaks,” “Increased” DE peaks, “Decreased” DE peaks, and those that do not overlap with any of the CTCF peaks identified in this study, respectively. (C) Enrichment of H3K9 methylation around SINE B3 transposons overlapping with “Increased” DE peaks. (D) Phylogenetic analysis of SINE B3 transposons. SINE B3 transposons overlapping with “Increased” DE peaks are color-coded according to their respective classes. (E) The distribution of Hi-C PC1 values in genomic regions where SINE B3 transposons overlap with “Increased” DE peaks. (F) Representative genomic regions where SINE B3 transposons overlap with “Increased” DE peaks. SINE B3 elements overlapping with Class 1 are predominantly found in strong A compartments, those overlapping with Class 2 are mainly in B compartments, and those in Class 3 are frequently located in intermediate regions between Class 1 and Class 2. (G) Epigenome profiles around IAPLTR2_Mm elements overlapping with “Increased” Class 5. In WT iMEFs, H3K9me3, regulated by SETDB1, is enriched. However, when H3K9 methylation is lost, H3K27me3 becomes enriched instead.











