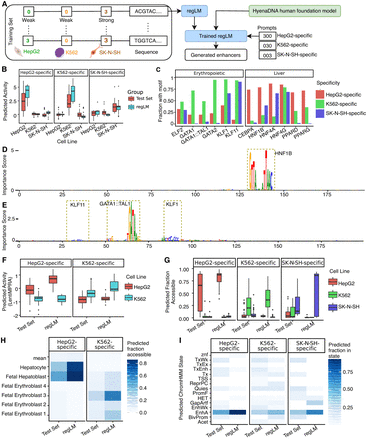
regLM generates synthetic cell type–specific human enhancers. (A) Schematic of the experiment. (B) Predicted activity of cell type–specific human enhancers generated by regLM, compared to real cell line-specific human enhancers from the test set, in three cell lines. (C) Fraction of regLM-generated enhancers containing selected cell type–specific TF motifs. (D) Sequence of a HepG2-specific regLM-generated enhancer. (E) Sequence of a K562-specific regLM-generated enhancer. Height is proportional to per-nucleotide importance scores from the independent regression model using ISM. Motifs with high importance are highlighted. (F) Predicted activity of real and regLM-generated cell type–specific enhancers, using a model trained on LentiMPRA data. (G) Predictions of a binary classification model trained on ATAC-seq from human cell lines, on real and regLM-generated cell type–specific enhancers. (H) Predictions of a binary classification model trained on pseudobulk snATAC-seq from 204 cell types, on real and regLM-generated cell type–specific enhancers. Color intensity represents the fraction of sequences in the group that were predicted to be accessible. “Mean” represents the average of all remaining cell types. (I) Predictions of a classification model trained to classify genomic DNA into chromatin states defined by the full-stack ChromHMM annotation, on real and regLM-generated cell type–specific enhancers. Color intensity represents the fraction of sequences in the group that were predicted to belong to the given state. (Acet) acetylations, (BivProm) bivalent promoter, (EnhA) enhancers, (EnhWk) weak enhancers, (GapArtf) assembly gaps and artifacts, (HET) heterochromatin, (PromF) Flanking promoter, (ReprPC) polycomb repressed, (Quies) quiescent, (TSS) transcription start site, (Tx) transcription, (TxWk) weak transcription, (TxEnh) transcribed enhancer, (TxEx) exon and transcription, (znf) ZNF genes.











