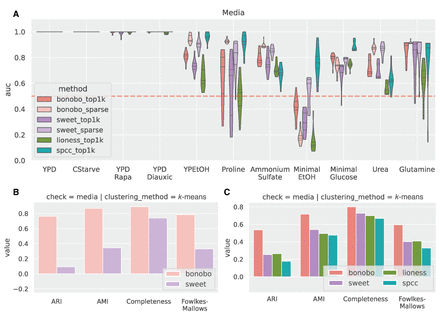
Comparison between methods for detecting media perturbations in yeast experiments. We used only the edges returned by the sparsified versions of SWEET and BONOBO and the 1000 strongest edges, on average, for each method (top1k labels). (A) Network similarity between perturbed yeast samples. For each sample, we computed Pearson correlation coefficients with all the other samples, which is always positive here, and estimated the ROC curve using a binary label that is one for the samples grown in the same media. For each comparison, we computed the AUROC (y-axis), grouping the samples by the growth medium (x-axis), and we compared the results between different methods. auc = 1 means that samples in the same media have the highest correlation values compared with samples in different media, auc = 0.5 (red dashed line) is the “random” performance, which means that samples with the same label are not more similar to each other than the rest. (B) Clustering performance on sparse BONOBO and SWEET. For all networks, we used k-means clustering and evaluated how well the resulting clusters captured growth media similarity using four different metrics: adjusted rand score (ARI), adjusted mutual info score (AMI), completeness, and Fowlkes–Mallows score. Greater values indicate that the clusters group samples grown in the same media. (C) Clustering performance on top1k networks using the same protocol as in B. Sparse networks, both for SWEET and BONOBO, outperform the naive “top1k” networks, with BONOBO outperforming SWEET. Although for some growth media, SPCC has a greater auc than BONOBO, BONOBO outperforms SPCC in the clustering test.











