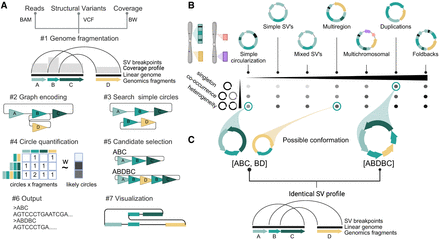
Decoil algorithm overview and an ecDNA ranking system based on its structural diversity. (A) Schematic of the Decoil algorithm depicting the major steps: (#1) genome fragmentation, (#2) graph encoding, (#3) search simple circles, (#4) circle quantification, (#5) candidate selection, (#6) output, and (#7) visualization. Step #7, visualization, is performed by the Decoil-viz module (see Methods). (B) ecDNA diversity. The x-axis displays the seven ecDNA topologies (e.g., simple circularization, multiregion, multichromosomal) with increasing computational complexity as defined in this paper. The y-axis displays different scenarios of ecDNA composition per sample, that is, singleton (presence of a single ecDNA structure), co-occurrence (presence of different ecDNA species, with nonoverlapping genomic regions), and heterogeneity (presence of different ecDNA species, with overlapping genomic regions). The gradient matrix depicts schematically the ecDNA reconstruction difficulty levels for the different scenarios (y-axis) and topologies (x-axis), which are addressed by Decoil algorithm. Light gray means low difficulty; black, increased difficulty. (C) Computational challenge formulation. The left panel displays a heterogeneity scenario, in which two different ecDNA elements (ABC, BD) share the genomic footprint (B fragment); the right panel displays a single large structure (ABDBC) containing interspersed-duplication rearrangement (B fragment duplicated on ecDNA). Both scenarios lead to the same SV breakpoint profile. To infer the likely conformation, we perform step #4 in A. Created with BioRender (https://www.biorender.com/).











