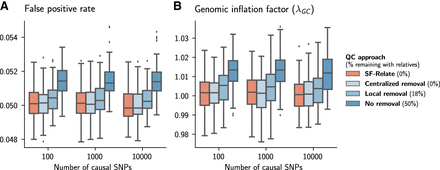
SF-Relate reduces false positives in multisite GWASs. We vary the number of causal SNPs used in simulating the phenotypes, and compare four quality-control (QC) approaches for excluding related individuals: (1) centralized removal (nonprivate), all relatives are removed from the pooled data set; (2) SF-Relate, relatives are removed using our secure approach; (3) local removal, each party filters relatives from its local data set independently; and (4) no removal, no relatives removed. Initially, 50% of samples have relatives, and local removal results in 18% of remaining samples still having a relative in the joint data set. We plot the fraction of significant loci (P-value < 0.05) on even numbered chromosomes that are designed to be noncausal in the simulation (A), and the genome inflation factor λGC in B. The filled boxes represent interquantile ranges of statistics across 100 simulated phenotypes. Although local removal of relatives helps reduce the confounding to some extent, SF-Relate significantly mitigates confounding, comparable to centrally coordinated sample removal.











