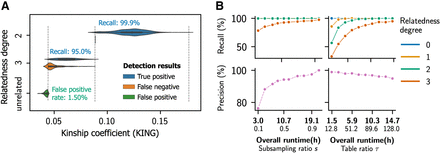
SF-Relate achieves higher accuracy for samples with closer kinship and enables a trade-off between accuracy and runtime. (A) We plot the distribution of kinship coefficients (KING) stratified by the (closest) relatedness degree of the relative pairs and by whether they were detected by SF-Relate as related. Misclassifications by SF-Relate are concentrated around kinship thresholds for different relatedness degrees, indicated by vertical dashed lines. (B) We vary the subsampling ratio (s) and the table ratio (τ) parameters in SF-Relate and report the resulting precision and recall for different relatedness degrees. For precision, only the overall metric for detecting third-degree or closer relatives is shown. By default, s = 0.7 and τ = 128. These parameters determine the trade-off between the runtime and accuracy of SF-Relate.











