
Mathematical and computational details of the PRiMeR framework. (A) Diagram illustrating the core MR assumptions, where genetic variants (g1, …, gS) influence exposure (e), which in turn affects outcome (o) with directional effect α. Additions unique to PRiMeR are highlighted in purple: a risk predictor is computed as a differentiable function 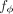 (parametrized by ϕ) of risk factors X. (B) Illustration of IVW regression, where genetic variant effects on outcome (βo) are regressed on the aggregate risk predictor (βe), accounting for their standard errors (so). (C) Main computations in PRiMeR, including computation of the risk predictor e(ϕ), the estimation of genetic effects on the risk predictor βe(ϕ), and the computation of the IVW regression loss. The function h(e(ϕ), G, F) returns marginal regression weights of each variant G:s on e(ϕ) accounting for covariates F. As all these steps are differentiable,
(parametrized by ϕ) of risk factors X. (B) Illustration of IVW regression, where genetic variant effects on outcome (βo) are regressed on the aggregate risk predictor (βe), accounting for their standard errors (so). (C) Main computations in PRiMeR, including computation of the risk predictor e(ϕ), the estimation of genetic effects on the risk predictor βe(ϕ), and the computation of the IVW regression loss. The function h(e(ϕ), G, F) returns marginal regression weights of each variant G:s on e(ϕ) accounting for covariates F. As all these steps are differentiable, 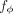 can be learned through gradient-based optimization of the IVW regression loss.
can be learned through gradient-based optimization of the IVW regression loss.











