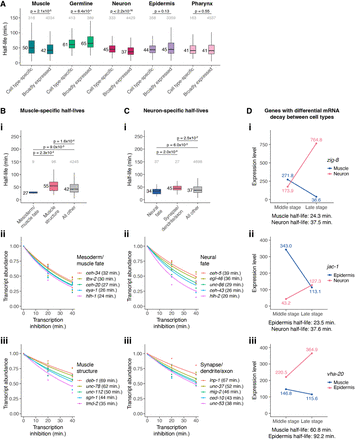
mRNA stability is correlated with cell type–specific functions. (A) Box plots showing the mRNA half-life distributions of cell type–specific and broadly expressed genes within muscle, germline, epidermis, neuron, and pharynx cells. Numbers to the left of the box plots are median half-lives within each group. Numbers above box plots are the number of genes within each group. The P-value comparing median half-lives was calculated using the Wilcoxon rank-sum test. Outliers are not shown: From left to right, 16, 125, 8, 28, one, 149, one, 69, five, and 120 genes within each group had mRNA half-lives >150 min. (B,i) Box plots showing the muscle-specific mRNA half-life distributions of mesoderm/muscle fate-specifying transcription factor genes, muscle structure genes, and all other genes. Numbers to the left of the box plots are median half-lives within each group. Numbers above box plots are the number of genes within each group. The P-value comparing median half-lives was calculated using the Wilcoxon rank-sum test. Outliers are not shown: From left to right, zero, four, and 137 genes within each group had mRNA half-lives >150 min. (ii,iii) Scatter plots of the normalized transcript abundance of the mesoderm/muscle fate-specifying transcription factor genes ceh-34, tbx-2, ceh-20, eya-1, and hlh-1 and the muscle structure genes deb-1, unc-78, unc-112, sgn-1, and tmd-2 throughout a 40 min transcription inhibition time course in muscle cells. Each point represents normalized transcript abundance from one of three biological replicates. (C,i) Box plots showing the neuron-specific mRNA half-life distributions of neural fate-specifying transcription factor genes, synapse-/dendrite-/axon-associated genes, and all other genes. Numbers to the left of the box plots are median half-lives within each group. Numbers above box plots are the number of genes within each group. The P-value comparing median half-lives was calculated using the Wilcoxon rank-sum test. Outliers are not shown: From left to right, zero, zero, and 70 genes within each group had mRNA half-lives >150 min. (ii,iii) Scatter plots of the normalized transcript abundance of the neural fate-specifying transcription factor genes ceh-5, egl-46, unc-86, ceh-43, and hlh-2 and the synapse-/dendrite-/axon-associated genes lnp-1, unc-37, mig-2, ced-10, and unc-53 throughout a 40 min transcription inhibition time course in neuronal cells. Each point represents normalized transcript abundance from one of three biological replicates. (D) Cell type–specific expression in the middle and late stages for genes with differential mRNA decay between muscle and neuron (i), epidermis and neuron (ii), and muscle and epidermis (iii). Cell type–specific expression in TPM was obtained from a C. elegans embryo single-cell atlas (Packer et al. 2019).











