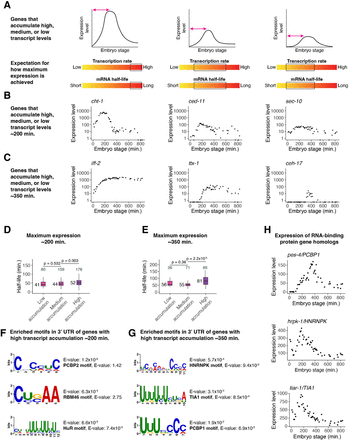
High transcript accumulation is associated with increased mRNA stability. (A) A schematic representation of genes that accumulate high, medium, or low transcript levels and the expected range of transcription and mRNA decay rates that could contribute to such accumulation. (B) Examples of genes that peak in expression ∼200 min after the four-cell stage with high, medium, and low transcript accumulation, from left to right. Expression data taken from a whole-embryo RNA sequencing time series (Hashimshony et al. 2015). (C) Examples of genes that peak in expression ∼350 min after the four-cell stage with high, medium, and low transcript accumulation, from left to right. Expression data taken from a whole-embryo RNA sequencing time series (Hashimshony et al. 2015). (D) Box plots showing the mRNA half-life distributions of genes that peak in expression ∼200 min after the four-cell stage to low, medium, and high transcript levels. Numbers to the left of the box plots are median half-lives within each group. Numbers above the box plots are the number of genes within each group. P-values comparing median half-lives were calculated using the Wilcoxon rank-sum test. Outliers are not shown: From left to right, one, zero, and seven genes within each group had mRNA half-lives >125 min. (E) Box plots showing the mRNA half-life distributions of genes that peak in expression ∼350 min after the four-cell stage with low, medium, and high transcript levels. Numbers to the left of the box plots are median half-lives within each group. Numbers above the box plots are the number of genes within each group. P-values comparing median half-lives were calculated using the Wilcoxon rank-sum test. Outliers are not shown: From left to right, zero, zero, and two genes within each group had mRNA half-lives >175 min. (F) MEME (Bailey et al. 2015) identified three motifs enriched in the 3′ UTRs of genes with high transcript accumulation compared with the 3′ UTRs of genes with low transcript accumulation ∼200 min after the four-cell stage. For genes with multiple 3′ UTR isoforms, the longest isoform was used. The best match of each de novo motif to known motifs in mammals (Ray et al. 2013) is noted. (G) MEME (Bailey et al. 2015) identified three motifs enriched in the 3′ UTRs of genes with high transcript accumulation compared with the 3′ UTRs of genes with low transcript accumulation ∼350 min after the four-cell stage. For genes with multiple 3′ UTR isoforms, the longest isoform was used. The best match of each de novo motif to known motifs in mammals (Ray et al. 2013) is noted. (H) The gene expression patterns for three putative RNA-binding protein genes in C. elegans from a whole-embryo RNA sequencing time series (Hashimshony et al. 2015). The genes are homologs of mammalian RNA-binding protein genes whose corresponding proteins are known to bind motifs similar to those discovered in F and G.











