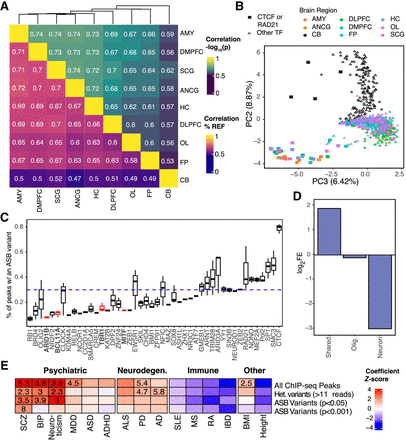
Neural-specific effects of ASB. (A) Correlation of ASB effects across brain regions for variants with a P-value < 0.001 in at least one brain region. (Top) Correlation of significance [−log10(P-value)]. (Bottom) Correlation of effect size (percentage of reads mapping to the reference allele [% REF]). (B) Dot plot showing PC2 and PC3 from a PCA of the −log10(P-values) for all variants with a P-value < 0.001 in at least one brain region. Color denotes the brain region. Shape denotes if the TF is CTCF/RAD21 or another TF in the data set. (C) Distribution of the percentage of peaks containing an ASB by TF. Dashed line indicates the average percentage. Only TFs with at least 15,000 peaks are shown. (D) Enrichment for ASB variants in cell type–specific snATAC-seq peaks. Neuron indicates that the peak was called in excitatory-only, inhibitory-only, or excitatory and inhibitory neurons. Shared peaks were identified in two or more cell types, excluding those shared between neuronal cell types. (E) Results from LDSC measuring heritability of ASB with GWAS traits grouped by disease type. Heatmap indicates coefficient Z-score from sLDSC. Feature-trait combinations with a Z-score significantly larger than zero (one-sided Z-test, P < 0.01) are indicated with a numeric value reporting the enrichment score. Abbreviations denote the following: (SCZ) schizophrenia, (BIP) bipolar disorder, (MMD) major depression disorder, (ASD) autism spectrum disorder, (ADHD) attention deficit hyperactivity disorder, (ALS) amyotrophic lateral sclerosis, (PD) Parkinson's disease, (AD) Alzheimer's disease, (SLE) systemic lupus erythematosus, (MS) multiple sclerosis, (RA) rheumatoid arthritis, (IBD) inflammatory bowel disease, (BMI) body mass index.











