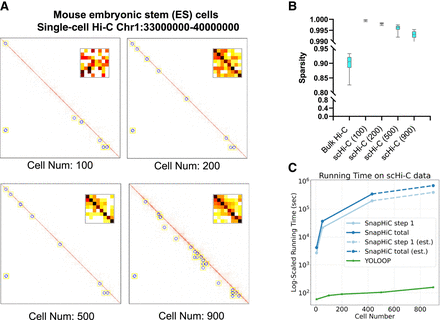
Fast adaptation of YOLOOP on single-cell data. (A) Superimposed single-cell contact matrices from 100 to 900 cells along with the detected loop patterns. (B) Sparsity comparison between bulk Hi-C and scHi-C data. (C) Prediction time comparison between YOLOOP and SnapHiC when increasing the imposed cell numbers. Time benchmarking for two methods both started from raw contact pair data. Estimations for SnapHiC's running time on 900 cells were interpolated owing to its time intensity.











