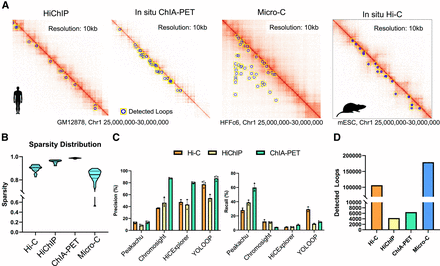
Fast adaptation of YOLOOP on multiple species and proximity ligation protocols. (A) YOLOOP is able to accurately capture loop anchors under multiple experimental protocols (ChIA-PET, HiChIP, Micro-C) in human and mouse cells. (B) Sparsity distribution for contact maps from various experiment protocols. (C) Statistical recovery rate and precision benchmarking YOLOOP against existing algorithms on Hi-C, ChIA-PET, and HiChIP GM12878 contact maps. (D) Enumeration of YOLOOP's predictions on contact maps from multiple protocols.











