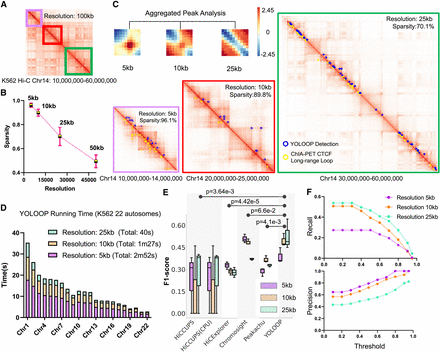
YOLOOP handling Hi-C maps on Homo sapiens K562 cells at multiple resolutions. (A) Sampled results on the K562 Hi-C contact matrix (Chr 14: 10,000,000–60,000,000). (B) Hi-C matrix sparsity statistics at 5 kb, 10 kb, and 25 kb resolution on Chromosome 14 of the H. sapiens K562 cell line. (C) Aggregated loop patterns and zoom-in detection results under multiple resolutions. (D) Running time for YOLOOP on 22 autosomes at multiple resolutions. (E) Statistical evaluation with F1-score benchmarking YOLOOP with existing methods across three different resolutions on the K562 cell line. Each box contains F1-scores distribution evaluated on all test chromosomes, with P-values calculated from the two-tailed t-test indicating the significance of the average differences across three resolutions. (F) The precision and recall curve of YOLOOP under various threshold selections.











