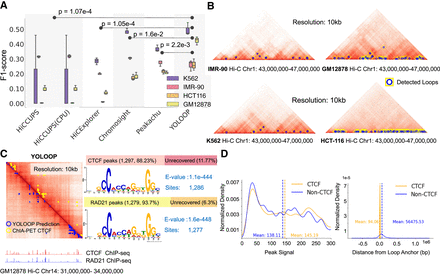
Application of YOLOOP on multiple cell lines and peak analysis. (A) Statistical evaluation with F1-score benchmarking YOLOOP against existing methods across four cell lines. Each box contains F1-score distributions evaluated on all test chromosomes. The P-values calculated from the two-tailed t-test indicates the significance of the average difference in F1-score between the YOLOOP and baseline methods across four cell lines. (B) Visualization of the detection results by YOLOOP on four cell lines. (C) Zoom-in detection results of YOLOOP on contact map of GM12878 cell for Chromosome 14 at 10 kb resolution. YOLOOP is able to recover the majority of the CTCF-enriched loops with a clear motif pattern (E-value denotes the statistical significance of the motif). The CTCF and RAD21 ChIP-seq signals in the anchors of the detected loops are shown under the contact map. (D, left) Density plots of the ChIP-seq signal value of the loops detected by YOLOOP against orthogonal CTCF binding sites in GM12878 cells. (Right) Density plots of the distance distribution for CTCF-supported loop anchors and non-CTCF-supported loop anchors.











