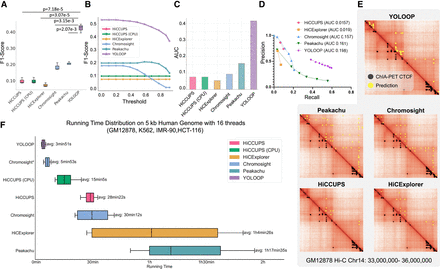
Performance and acceleration of YOLOOP in GM12878 Hi-C and comparison with baseline methods. (A) Statistical evaluation with F1-score, comparing YOLOOP against existing methods, with P-values calculated from the two-tailed t-test indicating the significance of the improvement. (B) Sensitivity evaluation. Each line represents the method's sensitivity to threshold selections. (C) Area under the curve (AUC) value of the sensitivity curve. (D) Precision-recall curve on GM12878 Hi-C with a 0.1 threshold gap with AUC reported across all benchmark methods. (For aesthetic purposes, we omitted connecting the (0,0) point.) (E) Visualization of orthogonal ChIA-PET CTCF binding sites and predicted loops by different methods. (F) Acceleration of YOLOOP compared with multiple baselines. The running time distribution is shown for four contact maps on human autosomes at 5 kb resolution, with five runs on each datum to increase statistical reliability. All multithreading-enabled algorithms were benchmarked with 16 CPU threads. Chromosight* indicates its running time after matrix balancing.











